I was shown this JOI problem by Jasonwei08 today. The problem statement boils down to:
Given an array $$$A$$$ of $$$2^L$$$ integers, $$$1 \leq L \leq 20$$$, process $$$1 \leq Q \leq 10^6$$$ queries. For each query, consisting of a length-$$$L$$$ pattern of either 0, 1, or ?, report the sum of all $$$A_j$$$ where the binary representation of $$$j$$$ matches the pattern.
The ? character acts as a wild card. For example, the pattern 1? matches both 10 and 11.
We will start with the following brute force approach:
Build a binary tree whose leaves correspond to all binary strings of length $$$L$$$, in lexicographic order. Each node's parent corresponds to the string obtained by deleting its last character, e.g. the parent of 11011 is 1101. We allow the tree's root node to be the empty string. It can be shown that this procedure yields a perfect binary tree.
We can visualize each query as a set of leaves in this tree. As a very simple example, let $$$L$$$ be $$$4$$$, and the query pattern be ?1?:
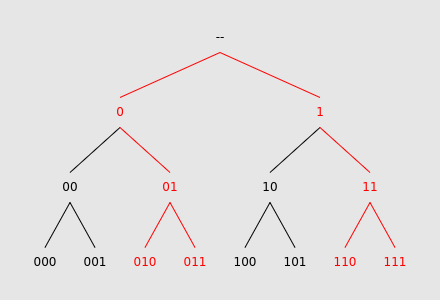
As expected, this pattern matches $$$010$$$, $$$011$$$, $$$110$$$, and $$$111$$$. Highlighted in red is the subtree whose leaves consist only of these matching strings, which we use to answer the query. By storing each array value $$$A_j$$$ within the leaf corresponding to $$$j$$$, and then performing a DFS on the subtree, we compute the answer to the query. Although this method may seem convoluted, it will allow for interesting optimizations later.
This approach yields a worst-case query complexity of $$$2^L$$$, which is unacceptable. How can we improve?
Let's look at the previous diagram again, and in particular the red subtree. Interestingly, it has a high degree of symmetry. Notice that the section rooted at node 0 has identical structure to the section rooted at node 1. If we think about how this subtree is defined, this property makes sense. Each ? in the query pattern essentially creates two sets of matching binary strings whose elements differ only at a single place value.
We can exploit this symmetry by "folding together" the tree at nodes 0 and 1, allowing two times fewer nodes to be traversed:
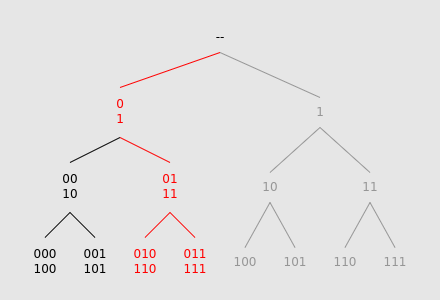
In order to support this "folding operation", we can store the tree's leaf values in an array. By cutting the array into two halves and summing their elements together, we obtain a new "leaf array" with half as many elements.
Notice that this "folding" step essentially reduces the original subtree into a line graph spanning from the root to the leaf level. This negates the need for a DFS. In fact, we can use this "leaf array" concept to implement the entire tree traversal:
As we descend down the subtree, we move to either the left child, right child, or both. In the first two cases, we can cut the leaf array in half, and select only one of them. In the third case, we "fold" as described before. Because this operation always results in a new array with exactly half the size of the previous one, our final array will contain only one element, whose value is identical to the answer of the query.
As an example, the query ?01? will be processed as follows:
Create the original leaf array
Cut the array in half, folding the two subarrays together
Cut the array in half, selecting the left one
Cut the array in half, selecting the right one
Cut the array in half and fold
By itself, this process still takes $$$2^L$$$ operations worst-case per query. However, we can improve this by leveraging cross-query similarities. In particular, if two different queries share the first couple of cut/fold operations, save the leaf array's state after such operations are applied, avoiding unnecessary recalculation.
This optimization turns out to be very useful. Because each cut/fold operation is linear with respect to leaf array size, the most costly operations run earliest, and are therefore most likely to be "memoized" by our optimization.
To implement this procedure, we collect all query patterns and build a trie. Then, we perform a depth-first search, which orders the queries optimally, avoiding all unnecessary cut/fold operations. Furthermore, if we are clever with the DFS order, we only need to store one leaf array of each size ($$$2^L, 2^{L-1}$$$, etc), which nicely bounds overall memory usage to $$$2^{L+1} - 1$$$.
Analyzing the time complexity of this algorithm is somewhat messy. Our worst-case scenario occurs when the query pattern trie has as many nodes as possible. Let us count the number of nodes at each depth level within such a worst-case trie. Each node can have at most three children, so the absolute maximum number of nodes at depth $$$i$$$ is bounded by $$$3^i$$$. Because we cannot have more nodes than the number of total queries, we restrict to $$$\min(3^i, Q)$$$. It can be shown that such a worst-case trie always exists.
A trie node at depth level $$$i$$$ operates on a leaf array of size $$$2^{L-i}$$$, so we can approximate the time complexity of the algorithm as:
To split up the $$$\min$$$, let $$$q = \left\lfloor \log_3 Q \right\rfloor$$$.
&= \sum_{i=0}^q 2^{L-i} 3^i + \sum_{q+1}^L 2^{L-i} Q \\
&= 2^L \sum_{i=0}^q 1.5^i + Q \left( 2^{L-q} - 1 \right) \\
&= 2^L \cdot 3 \cdot 1.5^q - 2^{L+1} + Q \frac{2^L}{2^q} - Q
\end{align} $$$
We can approximate $$$q$$$ as $$$\log_3 Q$$$ directly, and use change-of-base.
This is essentially on the order of $$$2^L Q^{0.369}$$$, which is very acceptable.
I have not seen this technique mentioned anywhere before. I think the original problem's intended solution was SOS DP, or at least that's how Jasonwei08 solved it. This cut/fold technique was definitely more work to implement than SOS, although probably not as complicated as it conceptually seems.
Generally, this technique can solve all sorts of pattern-matching problems, especially in situations where there are more than two types of characters and wildcards. The algorithm's time complexity scales nontrivially with respect to the number of character types, so I'm not sure how feasible it would be in extreme cases.
I do not expect this to be very useful in future contests, due to its extremely limited use case. However, I had a lot of fun discovering it, and I hope you enjoyed reading this blog. Let me know if I missed anything!









