Hello Codeforces,
CSES is a nice collection of classical CP problems which encourages you to learn a lot of basic and advanced concepts. Having editorials would help people to not get stuck on a problem for long. Here are my solutions to the tree section of the problem-set. Looking forward to read corrections/feedback/better solutions in the comment section. Happy coding!
Subordinates
This can be solved using basic DP approach in linear time.
$$$subordinates[u] = \sum\limits_{v:children[u]}^{} subordinates[v]$$$ where v is a child of u
Code#include<bits/stdc++.h>
using namespace std;
const int SZ = 200005;
int n, m, x;
vector<int> adj[SZ];
int S[SZ];
void dfs(int u,int l, int p)
{
S[u] = 1;
for (int v: adj[u]) {
if (v != p) {
dfs(v, l+1, u);
S[u] += S[v];
}
}
}
int main() {
cin >> n;
for(int i = 2; i <=n ; i++) {
cin >> x;
adj[i].push_back(x);
adj[x].push_back(i);
}
dfs(1, 0, 0);
for(int i = 1; i <= n ; i++) {
cout << S[i] - 1 << " ";
}
}
Tree Matching
This problem can be reduced to maximum bipartite matching problem but first we need to split the tree into a bipartite graph. One way to do that is to put all nodes at even depth in the first group and odd ones in the other group. Such a splitting will make sure that we do not have any edges within the same group.
Next, we can use Hopkraft Karp algorithm to find the maximum matching in the bipartite graph created.
Code#include<bits/stdc++.h>
using namespace std;
#define NIL 0
#define INF (1<<28)
const int SZ = 400005;
int x, u, v;
vector<int> adj_inp[SZ];
vector<int> adj[SZ];
int n, m, match[SZ], dist[SZ];
bool bfs() {
int i, u, v, len;
queue<int> Q;
for(i=1; i<=n; i++) {
if(match[i]==NIL) {
dist[i] = 0;
Q.push(i);
}
else dist[i] = INF;
}
dist[NIL] = INF;
while(!Q.empty()) {
u = Q.front(); Q.pop();
if(u!=NIL) {
for(int v: adj[u]) {
if(dist[match[v]]==INF) {
dist[match[v]] = dist[u] + 1;
Q.push(match[v]);
}
}
}
}
return (dist[NIL]!=INF);
}
bool dfs(int u) {
int i, v, len;
if(u!=NIL) {
for(int v: adj[u]) {
if(dist[match[v]]==dist[u]+1) {
if(dfs(match[v])) {
match[v] = u;
match[u] = v;
return true;
}
}
}
dist[u] = INF;
return false;
}
return true;
}
int hopcroft_karp_max_match() {
int matching = 0, i;
// match[] is assumed NIL for all vertex in adj
while(bfs())
for(i=1; i<=n; i++)
if(match[i]==NIL && dfs(i))
matching++;
return matching;
}
void make_bipartite_dfs(int u, int p, int l) {
if(l%2 and p != -1) {
adj[u].push_back(p+n);
adj[p+n].push_back(u);
}
for(int v: adj_inp[u]) {
if(v!=p) {
if(l%2) {
adj[u].push_back(v+n);
adj[v+n].push_back(u);
}
make_bipartite_dfs(v, u, l+1);
}
}
}
int main() {
cin >> n;
for(int i = 0; i < n-1; i++) {
cin >> u >> v;
adj_inp[u].push_back(v);
adj_inp[v].push_back(u);
}
make_bipartite_dfs(1, -1, 1);
cout << hopcroft_karp_max_match() << endl;
}
Tree Diameter
This is a classical problem having multiple solutions.
One easy to implement solution is using 2 Breadth First Searches (BFS). Start a BFS with a random node and store the last node encountered before search ends. This last node will definitely be one of the ends of the diameter (Why?). Now run a second BFS from this node and you will end on the other end of the diameter.
Code#include<bits/stdc++.h>
using namespace std;
#define SZ 200005
int n, m, x, y;
vector<int> adj[SZ];
pair<int,int> bfs(int src)
{
int d = 0;
queue<pair<int,int> > q;
q.push({src, 0});
vector<bool> vis(SZ, false);
pair<int,int> u;
while(!q.empty()) {
u = q.front();
vis[u.first] = true;
q.pop();
for(int v: adj[u.first])
if(!vis[v])
q.push({v, u.second + 1});
}
return u;
}
int main() {
cin >> n;
for(int i = 0; i < n-1; i++) {
cin >> x >> y;
adj[x].push_back(y);
adj[y].push_back(x);
}
pair<int,int> end1 = bfs(1);
pair<int,int> end2 = bfs(end1.first);
cout << end2.second << endl;
}
Tree Distances I
For each node, we want to calculate maximum distance to another node. Previously we saw that that if we start a BFS from any node, we end up on either of the diametric end. We can use this fact to efficiently compute the answer. Let's calculate distances of each node from both the ends of the diameter. Then maximum distance of each node can be calculated as:
max_distance[u] = max(distance_from_diametric_end1[u], distance_from_diametric_end2[u])
Code#include<bits/stdc++.h>
using namespace std;
#define SZ 200005
int n, m, k, x;
vector<int> adj[SZ];
vector<int> ans(SZ, -1);
int bfs(int src) {
int top;
queue<int> q;
vector<int> d(n+1, -1);
d[src] = 0;
ans[src] = max(ans[src], d[src]);
q.push(src);
while(!q.empty()) {
top = q.front();
q.pop();
for(int v: adj[top]) {
if(d[v] == -1) {
q.push(v);
d[v] = d[top] + 1;
ans[v] = max(ans[v], d[v]);
}
}
}
return top;
}
int main() {
int u, v;
cin >> n;
for(int i = 0; i < n-1; i++) {
cin >> u >> v;
adj[u].push_back(v);
adj[v].push_back(u);
}
int diam_end_1 = bfs(1);
int diam_end_2 = bfs(diam_end_1);
bfs(diam_end_2);
for(int i = 1; i <=n ; i++) {
cout << ans[i] << " ";
}
}
Tree Distances II
We can solve this problem using in-out DP on trees.
$$$in[u]:$$$ sum of distances from u to each node in subtree rooted at u
$$$out[u]:$$$ sum of distances from u to each node excluding the subtree rooted at u
Now, ans[u] = in[u] + out[u]
Calculating $$$in[u]$$$ is quite straightforward. Suppose we want to calculate the sum of distances starting at node u and ending at any node in subtree rooted at v. We can use the pre-calculated value for v and separately add the contribution created by edge $$$u\rightarrow v$$$. This extra quantity will be the subtree size of u. Then we can repeat the process for each child of u.
$$$in[u] = \sum\limits{}{}in[v] + sub[v]$$$ where v is a child of u
To calculate $$$out[u]$$$, first let's calculate the contribution of parent of u. We can use out[par] and add the difference created by the edge $$$u \rightarrow par_u$$$ which is n-sub[par]+1. Next, we add the contribution of each sibling of u: in[sib] + sub[sib]*2.
Finally we have the following formula:
$$$out[u] = out[par] + (n-sub[par] + 1) + (\sum\limits_{sib}{} in[sib] + sub[sib]*2) - (in[u] + sub[u]*2)$$$
Code#include<bits/stdc++.h>
using namespace std;
#define int long long int
#define SZ 200005
int n, m, k, x;
vector<int> adj[SZ];
int S[SZ];
int in[SZ];
int out[SZ];
void dfs_in(int u,int p)
{
S[u] = 1;
for (int v: adj[u]) {
if (v != p) {
dfs_in(v, u);
S[u] += S[v];
in[u] += in[v] + S[v];
}
}
}
void dfs_out(int u, int p) {
int store = 0;
for(int v: adj[u]) {
if(v != p)
store += in[v] + S[v]*2;
}
for (int v : adj[u]) {
if (v != p) {
out[v] = (out[u] + 1*(n-S[u]+1)) + (store - (in[v] + S[v] * 2));
dfs_out(v, u);
}
}
}
signed main() {
int u,v;
cin >> n;
for(int i = 0; i < n-1; i++) {
cin >> u >> v;
adj[u].push_back(v);
adj[v].push_back(u);
}
dfs_in(1, 0);
dfs_out(1, 0);
for(int i = 1; i <= n ; i++)
cout << in[i] + out[i] << " ";
}
Company Queries I
We can solve this problem using binary-lifting technique for finding ancestors in a tree.
For each node x given in a query, we just need to find the $$$k^{th}$$$ ancestor of a given node. We can initialise ans = x and keep lifting our ans for every $$$i^{th}$$$ bit set in k: ans = jump[i][ans] where $$$jump[i][j]$$$ holds $$$i^{th}$$$ ancestor of node $$$j$$$
Code#include<bits/stdc++.h>
using namespace std;
#define SZ 200005
int n, m, x,q,u,v,k;
int L[SZ];
vector<int> adj[SZ];
int jump[21][SZ];
void dfs(int u, int p, int l)
{
jump[0][u] = p;
L[u] = l;
for (int v: adj[u])
if(v != p)
dfs(v, u, l+1);
}
void preprocess_LCA()
{
dfs(1, -1, 0);
for (int i = 1; 1<<i <= n ; i++)
for (int j = 0; j <= n ; j++)
jump[i][j] = jump[i-1][jump[i-1][j]];
}
int main()
{
cin >> n >> q;
for(int i = 2; i <= n; i++) {
cin >> u;
adj[u].push_back(i);
adj[i].push_back(u);
}
preprocess_LCA();
while(q--) {
cin >> u >> k;
int boss = u;
for(int i=0; i<=20; i++)
if(k & (1<<i))
boss = jump[i][boss];
cout << (boss==0 ? -1 : boss) << endl;
}
}
Company Queries II
This is the classical problem of finding Lowest Common Ancestor which can be solved in multiple ways. One of the common ways is to use binary-lifting which requires $$$O(nlog(n))$$$ preprocessing and $$$O(logn)$$$ per query.
Code#include<bits/stdc++.h>
using namespace std;
#define SZ 200005
int n, m, x,q,u,v;
int L[SZ];
vector<int> adj[SZ];
int jump[21][SZ];
void dfs(int u, int p, int l)
{
jump[0][u] = p;
L[u] = l;
for (int v: adj[u])
if(v != p)
dfs(v, u, l+1);
}
void preprocess_LCA()
{
dfs(1, -1, 0);
for (int i = 1; 1<<i <= n ; i++)
for (int j = 0; j <= n ; j++)
jump[i][j] = jump[i-1][jump[i-1][j]];
}
int LCA(int p,int q)
{
if(L[p] < L[q])
swap(p, q);
int diff = L[p] - L[q];
for(int i = 20; i >= 0; i--)
if(diff & (1<<i))
p = jump[i][p];
if(p == q) return p;
for (int i = 20; i >= 0; i--) {
if (jump[i][p] != jump[i][q]) {
p = jump[i][p];
q = jump[i][q];
}
}
return jump[0][p];
}
int main()
{
cin >> n >> q;
for(int i = 2; i <= n; i++) {
cin >> u;
adj[u].push_back(i);
adj[i].push_back(u);
}
preprocess_LCA();
while(q--) {
cin >> u >> v;
cout << LCA(u, v) << endl;
}
}
Distance Queries
Distance between node u and v can be calculated as $$$depth[u] + depth[v] - 2*depth[LCA(u,v)]$$$.
LCA of (u,v) can be calculated using binary-lifting approach in $$$O(logn)$$$ per query.
Code#include<bits/stdc++.h>
using namespace std;
#define SZ 200005
int n, m, k, x;
int L[SZ]; //level-array
vector<int> adj[SZ];
int jump[21][SZ];
void dfs(int u, int p, int l) {
jump[0][u] = p;
L[u] = l;
for (int v: adj[u])
if(v != p)
dfs(v, u, l+1);
}
void preprocess_LCA() {
dfs(1, -1, 0);
for (int i = 1; 1<<i <= n ; i++)
for (int j = 0; j <= n ; j++)
jump[i][j] = jump[i-1][jump[i-1][j]];
}
int LCA(int p,int q) {
if(L[p] < L[q])
swap(p, q);
int diff = L[p] - L[q];
for(int i = 20; i >= 0; i--)
if(diff & (1<<i))
p = jump[i][p];
if(p == q) return p;
for (int i = 20; i >= 0; i--) {
if (jump[i][p] != jump[i][q]) {
p = jump[i][p];
q = jump[i][q];
}
}
return jump[0][p];
}
int main() {
int u, v, q, a, b;
cin >> n >> q;
for(int i = 0; i < n-1; i++) {
cin >> u >> v;
adj[u].push_back(v);
adj[v].push_back(u);
}
preprocess_LCA();
while(q--) {
cin >> a >> b;
int lca = LCA(a,b);
cout << L[a] + L[b] - 2*L[lca] << endl;
}
}
Counting Paths
This problem can be solved using prefix sum on trees.
For every given path $$$u \rightarrow v$$$, we do following changes to the prefix array.
prefix[u]++
prefix[v]++
prefix[lca]--
prefix[parent[lca]]--
Next, we run a subtree summation over entire tree which means every node now holds the number of paths that node is a part of. This method is analogous to range update and point query in arrays, when all updates are preceded by queries.
$$$prefix[u] = \sum\limits_{v:children[u]}^{} prefix[v]$$$
Code#include<bits/stdc++.h>
using namespace std;
#define SZ 200005
int n, m, k, x;
int L[SZ]; //L=level
vector<int> adj[SZ];
int jump[21][SZ];
vector<int> prefix(SZ, 0);
//calculates (parent,level) arrays
void dfs(int u, int p, int l) {
jump[0][u] = p;
L[u] = l;
for (int v: adj[u])
if(v != p)
dfs(v, u, l+1);
}
void preprocess_LCA() {
dfs(1, 0, 0);
for (int i = 1; 1<<i <= n ; i++)
for (int j = 0; j <= n ; j++)
jump[i][j] = jump[i-1][jump[i-1][j]];
}
int LCA(int p,int q) {
if(L[p] < L[q])
swap(p, q);
int diff = L[p] - L[q];
for(int i = 20; i >= 0; i--)
if(diff & (1<<i))
p = jump[i][p];
if(p == q) return p;
for (int i = 20; i >= 0; i--) {
if (jump[i][p] != jump[i][q]) {
p = jump[i][p];
q = jump[i][q];
}
}
return jump[0][p];
}
void dfs_prefix_sum(int u, int p) {
for(int v: adj[u]) {
if(v-p) {
dfs_prefix_sum(v, u);
prefix[u] += prefix[v];
}
}
}
int main()
{
int u, v;
cin >> n >> m;
for(int i = 0; i < n-1; i++) {
cin >> u >> v;
adj[u].push_back(v);
adj[v].push_back(u);
}
preprocess_LCA();
while (m--)
{
cin >> u >> v;
prefix[u]++;
prefix[v]++;
int lca = LCA(u, v);
prefix[lca]--;
prefix[jump[0][lca]]--;
}
dfs_prefix_sum(1, 0);
for(int i = 1; i <= n ; i++)
cout << prefix[i] << " ";
}
Subtree Queries
This problem can be solved by flattening the tree to an array and then building a fenwick tree over flattened array.
Once reduced to an array, the problem becomes same as point update and range query. We can flatten the tree by pushing nodes to the array in the order of visiting them during a DFS. This ensures the entire subtree of a particular node forms a contiguous subarray in the resultant flattened array. The range of indices corresponding to subtree of a node can also be pre-calculated using a timer in DFS.
Code#include<bits/stdc++.h>
using namespace std;
#define int long long int
#define SZ 200005
int n, m, k, x, q, s;
int timer;
int start[SZ];
int endd[SZ];
int value[SZ];
int bit[SZ];
vector<int> adj[SZ];
void update(int i, int delta) {
for(; i <= n; i += i&-i)
bit[i] += delta;
}
int query(int i) {
int sum = 0;
for(; i > 0; i -= i&-i)
sum += bit[i];
return sum;
}
void dfs(int u, int p) {
start[u] = ++timer;
update(timer, value[u]);
for(int v: adj[u]){
if(v!=p) {
dfs(v, u);
}
}
endd[u] = timer;
}
signed main() {
int type;
cin >> n >> q;
for(int i = 1; i <=n ; i++) {
cin >> value[i];
}
for(int i = 0; i < n-1; i++) {
int u,v;
cin >> u >> v;
adj[u].push_back(v);
adj[v].push_back(u);
}
dfs(1, 0);
while (q--)
{
cin >> type;
if(type == 1) {
cin >> s >> x;
update(start[s], -value[s]);
value[s] = x;
update(start[s], +value[s]);
}
else {
cin >> s;
cout << query(endd[s]) - query(start[s] - 1) << "\n";
}
}
}
Path Queries
This problem can be solved using Heavy-Light decomposition of trees.
First, we decompose the tree into chains using heavy-light scheme and then build a segment tree over each chain. A path from node u to v can seen as concatenation of these chains and each query can be answered by querying the segment trees corresponding to each of these chains on the path. At each node of segement tree, we store the sum of tree node values corresponding to that segment. Since heavy-light scheme ensures there can be at most $$$O(logn)$$$ chains, each query can be answered in $$$O(log^{2}n)$$$ time.
Similarly, each update can be performed in $$$O(logn)$$$ time as it requires update on a single chain (single segment tree) corresponding to the given node.
Code#include<bits/stdc++.h>
using namespace std;
#define int long long int
const int N = 200005;
struct node {
int l, r, mx;
node *left, *right;
node(int ll, int rr) :
l(ll), r(rr), mx(0), left(NULL), right(NULL) {}
}* root[N];
vector<int> chain[N];
vector<int> adj[N];
int sz[N], H[N], P[N], chainHd[N], pos[N],leadingEdge[N];
void update(node* p,int idx, int val) {
if(p->l == p->r) {
p->mx = val;
return;
}
int mid = (p->l + p->r)>>1;
update(idx<=mid ? p->left : p->right, idx, val);
p->mx = (p->left->mx + p->right->mx);
}
int query(node* p,int ql, int qr) {
if(qr < p->l or p->r < ql) return 0;
if(ql <= p->l and p->r <= qr) return p->mx;
return query(p->left,ql, qr) + query(p->right,ql,qr);
}
node* build(int n,int l, int r) {
node *p = new node(l, r);
if(l < r) {
int mid = (l + r)>>1;
p->left = build(n,l, mid);
p->right = build(n,mid+1, r);
}
p->mx= (l==r) ? leadingEdge[chain[n][l]] : (p->left->mx + p->right->mx);
return p;
}
void dfs_build(int n, int p, int h) {
P[n] = p;
H[n] = h;
sz[n] = 1;
for(int i=0; i<adj[n].size() ; i++) {
int v=adj[n][i];
if(v-p){
dfs_build(v, n, h+1);
sz[n] += sz[v];
}
}
}
void HLD(int n) {
chain[n].push_back(n);
for(int i = 0; i < chain[n].size();i++){
int v = chain[n][i];
chainHd[v] = n;
pos[v] = i;
for( int k=0;k<adj[v].size();k++) {
int j=adj[v][k];
if(j!=P[v])
(sz[j]<<1) >= sz[v] ? chain[n].push_back(j) : HLD(j);
}
}
root[n] = build(n,0, chain[n].size() -1);
}
int HLDquery(int u, int v) {
int ans = 0;
while(chainHd[u] != chainHd[v]) {
if(H[chainHd[u]] > H[chainHd[v]]) swap(u, v);
ans += query(root[chainHd[v]],0, pos[v]);
v = P[chainHd[v]];
}
if(pos[u] > pos[v])
swap(u, v);
ans += query(root[chainHd[u]],pos[u], pos[v]);
return ans;
}
signed main() {
ios::sync_with_stdio(0);cin.tie(0);
int q, n, a, b, c, k, ti, type;
cin >> n >> q;
for(int i = 1; i <= n ; i++) {
cin >> leadingEdge[i];
}
for(int i = 0; i < n-1; i++) {
cin >> a >> b;
adj[a].push_back(b);
adj[b].push_back(a);
}
dfs_build(1,1,0);
HLD(1);
while (q--)
{
int type;
cin >> type;
if(type==1)
{
cin >> k >> ti;
update(root[ chainHd[k] ], pos[k] , ti);
}
else
{
cin >> k;
cout << HLDquery(1,k) << endl;
}
}
}
Path Queries 2
This problem has similar solution as Path Queries. Instead of storing the sum of nodes in segment tree, we store the maximum of node values corresponding to that segment.
Code#include<bits/stdc++.h>
using namespace std;
#define lli long long int
#define vec vector
#define pb push_back
#define s(x) scanf("%d",&x)
#define all(x) x.begin(),x.end()
#define pii pair<int,int>
#define fi first
#define se second
#define SZ 200005
const int N = SZ;
struct node {
int l, r, mx;
node *left, *right;
node(int ll, int rr) :
l(ll), r(rr), mx(0), left(NULL), right(NULL) {}
};
/******** SEGTREE-ON EACH CHAIN ******/
void update(node* p,int idx, int val){
if(p->l == p->r) {
p->mx = val;
return;
}
int mid = (p->l + p->r)>>1;
update( (idx<=mid? p->left: p->right ), idx, val);
p->mx = max(p->left->mx, p->right->mx);
}
int query(node* p,int ql, int qr) {
if(qr < p->l or p->r < ql) return 0;
if(ql <= p->l and p->r <= qr) return p->mx;
return max(query(p->left,ql, qr), query(p->right,ql,qr));
}
node* build(int l, int r) {
node *p = new node(l, r);
if(l < r) {
int mid = (l + r)>>1;
p->left = build(l, mid);
p->right = build(mid+1, r);
}
return p;
}
/********* SETUP SUBSIZE,PARENT,HEIGHT ********/
vector<int> adj[N];
int n, q;
node* root[N];/**holds root of segtree of each chain**/
vector<int> chain[N];
int sz[N], H[N], P[N], chainIdx[N], pos[N];
void dfs_setup(int n, int p, int h) {
P[n] = p;
H[n] = h;
sz[n] = 1;
for(int i : adj[n]) {
if(i-p){
dfs_setup(i, n, h+1);
sz[n] += sz[i];
}
}
}
/************ HLD,HLD-QUERY***********************/
void HLD_build(int n) {
chain[n].pb(n);
for(int i = 0; i < chain[n].size();i++){
int v = chain[n][i];
chainIdx[v] = n;
pos[v] = i;
for(int j : adj[v])
if(j-P[v])
2*sz[j] >= sz[v] ? chain[n].pb(j) : HLD_build(j);
}
root[n] = build(0, chain[n].size() - 1);
}
int HLD_query(int u, int v) {
int ans = 0;
while(chainIdx[u] != chainIdx[v])
{
if(H[chainIdx[u]] > H[chainIdx[v]]) swap(u, v);
ans = max(ans, query(root[chainIdx[v]],0, pos[v]));
v = P[chainIdx[v]];
}
if(pos[u] > pos[v])
swap(u, v);
ans = max(ans,query(root[chainIdx[u]],pos[u], pos[v]));
return ans;
}
/******************************************************/
int main() {
ios::sync_with_stdio(0);
cin.tie(0);
cin >> n >> q;
int val[n];
for(int i=0; i<n; i++) {
cin >> val[i];
}
for(int i = 0;i < n-1;i++) {
int u, v;
cin >> u >> v;
u--;
v--;
adj[u].push_back(v);
adj[v].push_back(u);
}
dfs_setup(0, 0, 0);
HLD_build(0);
for(int i=0; i<n; i++) {
update(root[chainIdx[i]],pos[i], val[i]);
}
for(int i = 0;i < q;i++) {
int type;
cin >> type;
if(type == 1) {
int u, x;
cin >> u >> x;
u--;
update(root[chainIdx[u]],pos[u], x);
}else {
int u, v;
cin >> u >> v;
cout << HLD_query(u-1, v-1) << "\n";
}
}
}
Distinct Colors
This problem can be solved using Mo's algorithm on trees and can be reduced to this classical SPOJ Problem
Flatten the tree to an array using the same technique mentioned in solution of Subtree Queries. Then the subtree of each node will correspond to a contiguous subarray of flattened array. We can then use Mo's algorithm to answer each query in $$$O(\sqrt{n})$$$ time with $$$O(n\sqrt{n})$$$ preprocessing.
Code#include<bits/stdc++.h>
using namespace std;
#define SZ 200005
int n, m, k, x, SQRT, ans, u, v;
vector<int> arr;
vector<int> adj[SZ];
int col[SZ];
int start[SZ];
int finish[SZ];
int t = -1;
int freq[SZ];
int query_ans[SZ];
struct node{
int L,R,i;
} query[SZ];
void dfs_flatten(int u, int p) {
start[u] = ++t;
arr.push_back(col[u]);
for(int v: adj[u]) {
if(v-p)
dfs_flatten(v, u);
}
finish[u] = t;
}
void map_colors(int col[]) {
map<int, int> prev_to_new;
vector<int> copy_col(col+1, col+1+n);
sort(copy_col.begin(), copy_col.end());
for(int i = 0; i < n; i++) {
prev_to_new[copy_col[i]] = i;
}
for(int i = 1; i <= n ; i++) {
col[i] = prev_to_new[col[i]];
}
}
bool cmp(node a,node b) {
if (a.L/SQRT != b.L/SQRT)
return a.L < b.L;
return a.R < b.R;
}
void add(int pos) {
freq[arr[pos]]++;
if(freq[arr[pos]]==1)
ans++;
}
void remove(int pos) {
freq[arr[pos]]--;
if(freq[arr[pos]]==0)
ans--;
}
void mos_algorithm() {
sort(query+1, query+1+n, cmp);
int currentL = 0, currentR = 0, L, R;
add(0);
for(int i=1; i<=n; i++){
L=query[i].L;
R=query[i].R;
while(currentL < L)
remove(currentL++);
while(currentL > L)
add(--currentL);
while(currentR < R)
add(++currentR);
while(currentR > R)
remove(currentR--);
query_ans[query[i].i]=ans;
}
}
int main() {
cin >> n;
SQRT=floor(sqrt(n));
for(int i = 1; i <= n; i++) {
cin >> col[i];
}
map_colors(col);
for(int i = 0; i < n-1; i++) {
cin >> u >> v;
adj[u].push_back(v);
adj[v].push_back(u);
}
dfs_flatten(1, 0);
for(int i = 1; i <= n; i++) {
query[i].L = start[i];
query[i].R = finish[i];
query[i].i = i;
}
mos_algorithm();
for(int i=1; i<=n; i++)
cout << query_ans[i] << " ";
}
Alternate solution using small-to-large technique#include<bits/stdc++.h>
using namespace std;
#define SZ 200005
int n, m, k, u, v;
vector<int> adj[SZ];
int ans[SZ];
int col[SZ];
set<int> dfs(int u, int p) {
set<int> uniq;
uniq.insert(col[u]);
for(int v: adj[u]) {
if(v!=p) {
set<int> child = dfs(v, u);
if(child.size() > uniq.size()) swap(child, uniq);
for(int color: child) {
uniq.insert(color);
}
}
}
ans[u] = uniq.size();
return uniq;
}
int main() {
cin >> n;
for(int i = 1; i <= n; i++)
cin >> col[i];
for(int i = 0; i < n-1; i++) {
cin >> u >> v;
adj[u].push_back(v);
adj[v].push_back(u);
}
dfs(1, 0);
for(int i=1; i<=n; i++)
cout << ans[i] << " ";
}
Finding a Centroid
We can precompute subtree sizes in a single DFS. We can recursively search for the centroid with one more DFS.
Code#include<bits/stdc++.h>
using namespace std;
#define SZ 200005
vector<int> adj[SZ];
int sub[SZ], n;
void dfs_sub(int u, int p) {
sub[u]=1;
for(int v: adj[u]) {
if(v!=p) {
dfs_sub(v, u);
sub[u] += sub[v];
}
}
}
int centroid(int u, int p) {
for(int v: adj[u]) {
if(v!=p) {
if(sub[v]>n/2) {
return centroid(v, u);
}
}
}
return u;
}
int main() {
ios::sync_with_stdio(0);
int u, v;
cin >> n;
for(int i=1; i<=n-1; i++) {
cin >> u >> v;
adj[u].push_back(v);
adj[v].push_back(u);
}
dfs_sub(1, -1);
cout << centroid(1, -1);
}
Fixed-Length Paths 1
Prerequisite: Centroid Decomposition of a tree
Hint: At every step of centroid decomposition, we try to calculate the number of desirable paths passing through the root(centroid). Since all nodes will eventually become centroid of a subtree, we claim all possible paths are considered. While processing a particular node, we can count the number of nodes at particular depth with cnt[d]++ and use this information to compute our result efficiently.
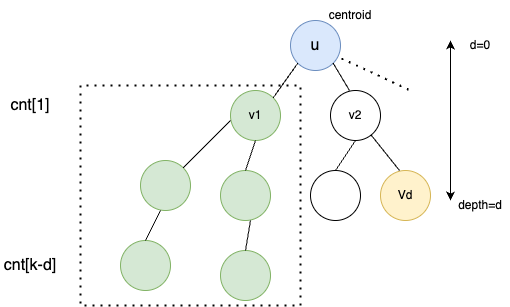
Explanation: Consider the diagram aboce where we are currently solving for tree rooted at $$$u$$$.
Assuming we have already processed $$$v_{1}$$$ and its subtree. We have updated the cnt array accordingly. While processing node $$$V_{d}$$$, we know it can create $$$cnt[k-d]$$$ new paths of length $$$k$$$. Hence, we add $$$ans += cnt[k-d]$$$.
Once $$$ans$$$ is updated, we can process the subtree of $$$v_{2}$$$ and update $$$cnt$$$ which can be used while processing other children of $$$u$$$.
After processing the whole subtree of $$$u$$$, we decompose the tree by removing the centroid $$$u$$$ and repeat the same process for disjoint trees rooted at $$$v_{1},v_{2}, v_{3}$$$ and so on.
Complexity: Centroid tree has a height of $$$log(n)$$$. Each horizontal layer of tree takes an amortised time of $$$O(n)$$$ as explained above. Thus, total complexity becomes $$$O(n*logn)$$$.
Code#include<bits/stdc++.h>
using namespace std;
#define SZ 200005
vector<int> adj[SZ];
int sub[SZ], cnt[SZ], n, k, max_d;
long long ans;
bool vis[SZ];
void dfs_sub(int u, int p) {
sub[u]=1;
for(int v: adj[u]) {
if(!vis[v] && v!=p) {
dfs_sub(v, u);
sub[u] += sub[v];
}
}
}
int find_centroid(int u, int p, int treeSz) {
for(int v: adj[u]) {
if(!vis[v] && v!=p && sub[v]>treeSz) {
return find_centroid(v, u, treeSz);
}
}
return u;
}
void dfs(int u, int p, int d, bool isCalculate) {
if(d>k) return;
if(isCalculate) {
ans += cnt[k-d];
}
else {
cnt[d]++;
}
max_d = max(max_d, d);
for(int v: adj[u]) {
if(!vis[v] && v!=p) {
dfs(v, u, d+1, isCalculate);
}
}
}
void centroid_decompose(int u) {
dfs_sub(u, -1);
int centroid = find_centroid(u, -1, sub[u]>>1);
vis[centroid] = true;
cnt[0] = 1;
max_d = 0;
for(int v: adj[centroid]) {
if(!vis[v]) {
dfs(v, centroid, 1, true);
dfs(v, centroid, 1, false);
}
}
fill(cnt, cnt+max_d+10, 0);
for(int v: adj[centroid]) {
if(!vis[v]) {
centroid_decompose(v);
}
}
}
int main () {
ios::sync_with_stdio(0);
cin.tie(0);
cin >> n >> k;
for (int i = 1; i < n; i++) {
int u, v;
cin >> u >> v;
adj[u].push_back(v);
adj[v].push_back(u);
}
centroid_decompose(1);
cout << ans;
}
Fixed-Length Paths 2
Approach 1: you can use centroid decomposition here as well but this time sum of all nodes in depth range $$$[k1-d,k2-d]$$$ must be added to $$$ans$$$. This is because all of these nodes will form a path length in range $$$[k1, k2]$$$. This can be done by using a Fenwick tree to calculate range sum efficiently. Although this works in $$$O(n*log^{2}n)$$$, this approach fails CSES time limits.
Code#include<bits/stdc++.h>
using namespace std;
#define SZ 200005
#define ll long long
vector<int> adj[SZ];
int sub[SZ], n, k1, k2, max_d;
long long ans;
bool vis[SZ];
/**-- fenwick tree --**/
int bit[200005];
void update(int i, ll delta) {
i++; // hack to allow 0-based indexing in fenwick tree
for(; i <= n; i += i&-i)
bit[i] += delta;
}
int query(int i, int i2) {
i2++;
int sum=0;
for(; i2 > 0; i2 -= i2&-i2)
sum += bit[i2];
for(; i > 0; i -= i&-i)
sum -= bit[i];
return sum;
}
/**-------------------**/
void dfs_sub(int u, int p) {
sub[u]=1;
for(int v: adj[u]) {
if(!vis[v] && v!=p) {
dfs_sub(v, u);
sub[u] += sub[v];
}
}
}
int find_centroid(int u, int p, int treeSz) {
for(int v: adj[u]) {
if(!vis[v] && v!=p && sub[v]>=treeSz) {
return find_centroid(v, u, treeSz);
}
}
return u;
}
void dfs(int u, int p, int d, bool isCalculate) {
if(d>k2) return;
if(isCalculate)
ans += (query(max(0,k1-d), k2-d));
else
update(d, +1);
max_d = max(max_d, d);
for(int v: adj[u]) {
if(!vis[v] && v!=p) {
dfs(v, u, d+1, isCalculate);
}
}
}
void centroid_decompose(int u) {
dfs_sub(u, -1);
int centroid = find_centroid(u, -1, sub[u]>>1);
vis[centroid] = true;
max_d = 0;
for(int v: adj[centroid]) {
if(!vis[v]) {
dfs(v, centroid, 1, true);
dfs(v, centroid, 1, false);
}
}
for(int i=1;i<=max_d;i++) {
update(i, -query(i,i));
}
for(int v: adj[centroid]) {
if(!vis[v])
centroid_decompose(v);
}
}
int main () {
cin.tie(0);
ios::sync_with_stdio(0);
cin >> n >> k1 >> k2;
if(k2-k1==n-1) {
cout << (1LL*n*(n-1))/2;
return 0;
}
for (int i = 1; i < n; i++) {
int u, v;
cin >> u >> v;
adj[u].push_back(v);
adj[v].push_back(u);
}
update(0, +1);
centroid_decompose(1);
cout << ans;
return 0;
}
Approach 2: Prerequisite: Small-To-Large Merging
Instead of using centroid decomposition, we use small-to-large technique to reduce operations. And instead of fenwick tree we use suffix sum arrays to compute range sums efficiently.
For a particular node $$$u$$$ and a particular child $$$v$$$ of $$$u$$$, we maintain suffix sum of count of nodes present at a depth d from u.
$$$suf[i] =$$$ count of nodes having depth in range $$$[i, inf)
$$$
Note that we can combine 2 suffix sum arrays efficiently in $$$O(min(|a|, |b|)$$$ using small-to-large merging technique.
For eg. $$$a=[a_{1},a_{2},a_{3}]
$$$
$$$b=[b_{1},b_{2}] $$$
can be combined as $$$[a_{1}+b_{1}, a_{2}+b_{2}, a_{3}]$$$
As stated in approach 1, for every node we want to know count of nodes in depth range $$$[k1-d, k2-d]$$$. We can calculate this contribution whenever we are merging smaller suffix sum array to larger one.
Source: https://usaco.guide/problems/cses-2081-fixed-length-paths-ii/solution
Code#include<bits/stdc++.h>
using namespace std;
#define SZ 200005
#define ll long long
vector<int> adj[SZ];
int n, k1, k2;
long long ans;
int suf(deque<int> &suf, int idx) {
if(idx<0) return suf[0];
if(idx>suf.size()-1) return 0;
return suf[idx];
}
/* suf[i] -> sum of nodes having depth in range [i, inf) */
void mergeSuffixes(deque<int> &sa, deque<int> &sb) {
if(sa.size() < sb.size()) swap(sa, sb);
for(int i=0; i<sb.size(); i++)
ans += 1LL * (suf(sb,i) - suf(sb, i+1)) * (suf(sa, k1-i) - suf(sa, k2-i+1));
for(int i=0; i<sb.size(); i++)
sa[i] += sb[i];
}
deque<int> dfs(int u, int p) {
deque<int> suf_parent{1};
for(int v: adj[u]) {
if(v!=p) {
deque<int> suf_child = dfs(v, u);
suf_child.push_front(suf_child.front()); // bcoz depth of child[0] should correspond to parent[1]
mergeSuffixes(suf_parent, suf_child);
}
}
return suf_parent;
}
int main () {
cin.tie(0)->sync_with_stdio(0);
cin >> n >> k1 >> k2;
for (int i = 1; i < n; i++) {
int u, v;
cin >> u >> v;
adj[u].push_back(v);
adj[v].push_back(u);
}
dfs(1,-1);
cout << ans;
return 0;
}
End Notes











